복잡한 세포 Assay의 3D 오가노이드 및 자동화[팟캐스트]
As we enter the era of sophisticated drug discovery with gene therapy and personalized medicine, we need to be prepared to study complex diseases, assess the therapeutic effect of drugs and identify adverse effects that can pose risks to patient health. Unfortunately, the current preclinical methods, such as animal models or 2D cell cultures, are inadequate. Because the physical and chemical properties of these models do not represent the human condition, preclinical drug evaluation does not translate to clinical success. That’s why the development of 3D cell models, such as organoids, can be a huge milestone for improving the evaluation of drug efficacy and safety.
Dr. Oksana Sirenko is the senior manager of assay development at Molecular Devices, working on the development of complex cell-based models for 3D biology, as well as high-content imaging and assay automation.

In this podcast excerpt, Senior scientist Oksana discusses the advantages of 3D cell models while addressing challenges in 3D cell imaging, such as image quality, high throughput, automation, and analysis.
Table of Contents
1. Why are 3D cell models and 3D organoids so useful in disease research and drug screening?
The main problem in current disease research and drug development is that only about 3% of developed drugs make it to the clinic. The majority of drugs fail in clinical trials because of a lack of efficacy or unwanted toxicity problems. Better assay systems and disease models are needed to facilitate drug discovery and better predict success in the clinic.
Today, biology is shifting toward increased complexity for assays and models that can be used for drug discovery and development. 3D models are believed to bridge the gap between traditional cell-based models, and tissues and organs. 3D models, which include spheroids, organoids, and organ-on-a-chip, present a variety of human cell types, such as liver, immune cells, cardiac cells, and fibroblasts. Also, they can mimic the morphology of human tissue types, such as 3D tumor growth, crypts in intestinal organoids, neural tubes, or the flow of liquids. Finally, they represent at least some aspects of tissue functionality, from the metabolic activity of the liver to the beating of cardiac organoids to spikes of neuronal activity in brain organoids. This greater complexity and sophistication allow us to mimic processes in tissues, interactions between cell types, responses to drugs, toxicity effects, and processes of drug penetration into the tissue.
https://share.vidyard.com/watch/savfn77oaeR6fQfcfJwZ54
Cerebral organoids show organization reminiscent of a developing brain.
2. Why does the complexity of 3D models present a hurdle/challenge for researchers?
Traditional 2D cell assays are easier to work with, but 3D assays have greater predictability and allow the generation of more biologically relevant data. However, despite the increasing interest in 3D research, the wide adoption of assays is limited by technical hurdles and assay complexity, which leads to higher costs, lower throughput, and a lack of reproducibility. Greater complexity presents challenges, but opportunities for instrument development and automation development would enable scientists to run 3D assays with greater throughput and accuracy.
3. Can you outline a typical workflow for 3D organoid development and analysis?
The typical workflow for organoids assays contains a number of steps, and this process is typically much longer than 2D workflow steps.
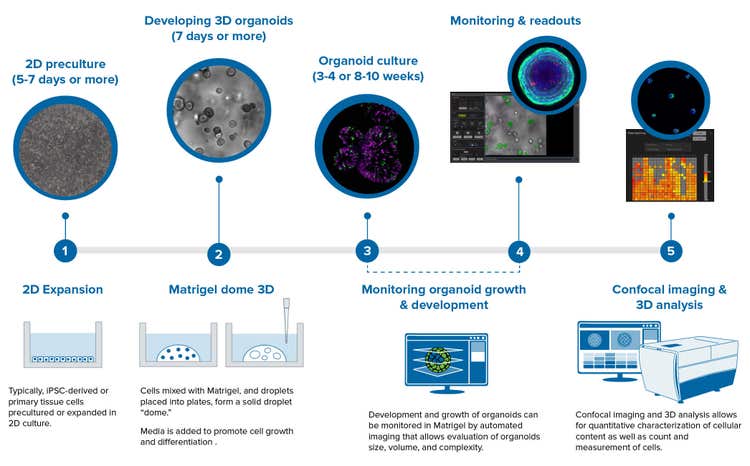
3D organoids can be derived from primary cells like intestinal organoids or induced pluripotent stem cells (iPSCs), e.g., neuronal or cardiac organoids. The workflow may start from 2D pre-culture or expansion of iPSC cells, followed by the differentiation step. After that, cells are mixed with Matrigel, and typically developed inside Matrigel domes, which also may include passaging and expansion. Intestinal organoids, colorectal, pancreatic, and liver are typically developed using this Matrigel dome step.
Alternatively, some other organoid types do not require Matrigel but instead are developed in low attachment plates (e.g., cardiac organoids).
Development of organoids takes from a few days to several weeks. Some protocols even require a few months. This is a very tedious process and will greatly benefit from automation.
Finally, the endpoint assay, whether it is a drug treatment, viral infectivity assay, or toxicity assessment, is set up in a multi-well format, with 96 or 384-well plates.
Next, cells are treated with drugs and processed for read-outs, which may include ATP assays, cell death assays, high-content imaging, or calcium oscillation.
4. Can you tell me a bit about how you apply 3D cell model workflows specifically to your research?
We are focusing on the development of automation protocols for automated cell cultures, as well as automated imaging and image analysis for complex 3D workflows. Recently, we developed and ran automated screening assays for finding more efficient anti-cancer drugs for triple-negative breast cancer. We used patient-derived cancer organoids representing a drug-resistant disease phenotype, and we applied automation to culture 3D organoids, simulate drug intervention, and run end-point assays for identifying the compounds that kill tumor cells. We tested a library of compounds and found several candidates that had greater efficacy than current standard drugs.
5. Can you explain how to automate the workflow for the development and analysis of organoids?
We created an automated workcell at Molecular Devices that combines several instruments in one complex system. It includes a Beckman Biomek automated liquid handler, a LiCONiC automated incubator, our ImageXpress HT.ai high-content imaging system, our SpectraMax plate reader and AquaMax washer, and a Bionex automated centrifuge. All the components are connected by a collaborative robot, PreciseFlex 400 that can move plates from one instrument to another at desired time points, while scheduling software ensures that all system elements work together seamlessly. Each instrument has multiple protocols designed for different steps, including feeding the cells and organoid plating, which can be called out by the scheduler at indicated times.

The Organoid Innovation Center at Molecular Devices combines cutting-edge technologies with novel 3D biology methods to address key challenges of scaling complex 3D biology.
Imaging methods are another exciting area of technology for organoid research. To image organoids or organ-on-a-chip, we need to use advanced optics. The ImageXpress high-content imaging system has several advantages for 3D samples:
Powerful lasers and confocal optics allow us to take the Z-stack of multiple images starting from the bottom and going up with steps like 5-10 microns Confocal optics allow us to reject the light that is out of focus so that we can get sharper images throughout organoids and Matrigel.
Next, our MetaXpress image analysis software analyzes the images in each 2D slice and converts data into 3D space. You can get multiple measurements to characterize organoids, cells, or subcellular organelles. These measurements help define cell counts, intensities, volumes, area, distances, and more, allowing us to monitor and quantitate changes in morphology, cell content, and viability. We also have machine learning elements, where users can train software to recognize objects, and features, to provide more efficient and insightful analysis.
6. How can automation aid in the research of complex systems specifically?
Automating would reduce labor and repetitive tasks like feeding cells every day or every second day for 2 months. Also, it will help to ramp up the research with higher throughput. For example, instead of studying 3 cell lines or 5 mutations, automation would allow you to test 50 cell lines to study 100 mutations.
High-content imaging powered by machine learning algorithms will allow to observe and characterize a variety of changes in organoids and cells, providing multiple readouts and yielding a valuable set of information about cell growth, differentiation, cell cycle, death, apoptosis, gene expression, or activation of signaling pathways.
7. How will you use these systems again in your future research?
In addition to cancer biology studies, we are actively developing other workflows, including but not limited to intestinal organoids, stem cell workflow, cardiac organoids, and more.
8. How will the automation of 3D organoid analysis evolve in the future?
We believe, as biology evolves and the complexity of assays increases, automation will be increasingly important for better understanding disease mechanisms, accelerating drug discovery, and eventually finding better ways to treat diseases.
By developing new and more advanced technologies and instruments, we believe we will further contribute to the progress of life sciences.
Understanding the basic principles behind 3D organoids - and the current bottlenecks - is crucial to the successful development and utilization of these advanced models for drug discovery.
Listen to the full podcast
If you'd like to learn more, please enjoy the full podcast, “Complex assays with Ian Shoemaker, Beckman Coulter Life Sciences & Dr Oksana Sirenko, Molecular Device."
Drug Target Review · Episode 14 - Ian Shoemaker, Beckman Coulter & Dr Oksana Sirenko, Molecular Devices
Here they discuss a comprehensive outlook on the science of organoid research – discover how organoid models are developed, used within our experts’ research, and how the automation of 3D organoid analysis will evolve in the future, plus much more!